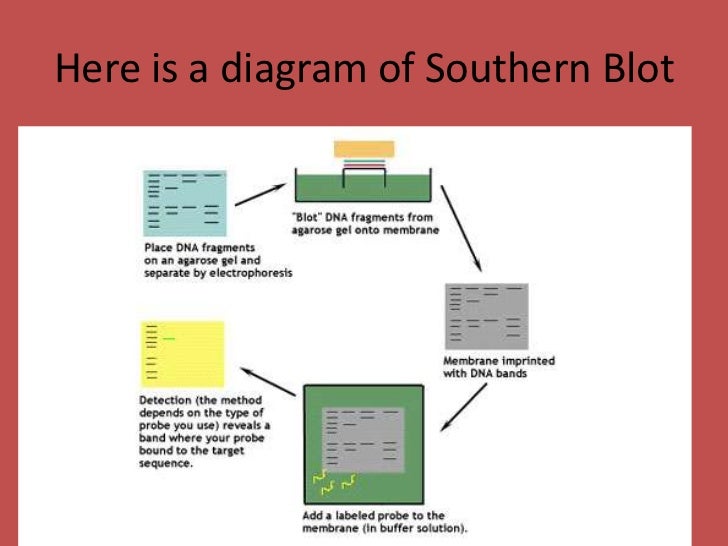
Suddenly, scientists could detect a segment of DNA without purifying it from the rest of the genome. This procedure generated a high-resolution picture of the DNA bands that held sequences of interest. After applying radiolabeled RNA to the membrane and washing off all of it that didn’t stick to matching DNA sequences, Southern exposed the membrane to X-ray film. During pilot experiments, he realized that the trick would be to soak the DNA fragments out of the gel by forcing liquid to flow through the gel onto the nitrocellulose he could accomplish this task by piling dry filter paper on top of the nitrocellulose, which would draw the liquid that would carry the DNA. 
However, he needed a way to transport the DNA. If he could move DNA fragments from the gel to a membrane made of nitrocellulose, which grabs and clings to DNA, he knew he could then bind radioactively labeled RNA to the trapped DNA because that method was well established. The tedium and labor involved in such a scheme spurred Southern to think of a better way. Southern realized that he could accomplish his task by brute force: carving the gel into small horizontal slabs washing the DNA out of each gel slice attaching every portion to a separate filter fishing for the particular DNA with a piece of matching, radioactively tagged RNA that would bind to it and then measuring the amount of bound radioactivity. Finding a single piece of DNA that carried a specific sequence was hopeless. For organisms with large genomes, however, this procedure generated a smear of DNA because of the millions of fragments. The pieces would migrate at different rates, depending on size. They could then separate the resulting pieces by loading the collection onto an agarose gel and applying an electric current. Scientists knew that they could chop up DNA using restriction enzymes, proteins that cut DNA at particular sequences. In the mid-1970s, Ed Southern (at the Medical Research Council Mammalian Genome Unit in Edinburgh) wanted to develop a method that would pinpoint a particular gene amidst the more than a billion building blocks - or basepairs - that compose the frog Xenopus laevis genome. This situation severely restricted efforts to define genetic differences that characterize species, individuals, and specific cell types, thus hampering the study of subjects as diverse as evolution and the physiological characteristics of distinct tissues. Until the mid-1970s, the ability to locate most genes or sequences of interest on the chromosomes of complex organisms was nearly impossible. Technology has always defined the strength of genetic analysis. Its ability to establish family relationships as well as individual identity has helped solve crimes, settle paternity and immigration disputes, establish the bases of inherited diseases, enhance transplantation biology, save endangered species, establish human origins and migrations, and advance countless other beneficial endeavors. Using this technology, Alec Jeffreys devised 'genetic fingerprinting', a way to distinguish every person from every other person, except an identical twin. Suddenly scientists could study genetic variation in detail and decipher gene structures. By inventing a method for detecting specific DNA sequences amidst the huge genomes of complex organisms, Edwin Southern infused genetic analysis with tremendous power. The 2005 Albert Lasker Award for Clinical Medical Research honors two scientists who revolutionized human genetics and forensic diagnostics.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
